Training and Evaluation
With the feature matrix built and preprocessed, this page covers the final step of the pipeline: splitting the data, training three classification models, and comparing their predictive performance with AUC.
Load Preprocessed Data
The preprocessing step saved the final training and test splits to CSV. Load them here:
using CSV, DataFrames
using CategoricalArrays: categorical
using MLJ
train_df = CSV.read(joinpath("output", "train.csv"), DataFrame)
test_df = CSV.read(joinpath("output", "test.csv"), DataFrame)
# MLJ classifiers require a CategoricalVector of strings for the target
train_df.outcome = categorical(string.(train_df.outcome); levels = ["0", "1"])
test_df.outcome = categorical(string.(test_df.outcome); levels = ["0", "1"])
y_train = train_df.outcome
X_train = select(train_df, Not(:outcome))
y_test = test_df.outcome
X_test = select(test_df, Not(:outcome))The levels = ["0", "1"] call ensures both the train and test vectors share the same encoding - important for models that inspect the level order internally.
Train-Test Split
The 80/20 split is performed during preprocessing (see preprocessing.jl) with a fixed random seed so the partition is reproducible:
# Done in preprocessing.jl:
# train, test = partition(df, 0.8; shuffle = true, rng = 42)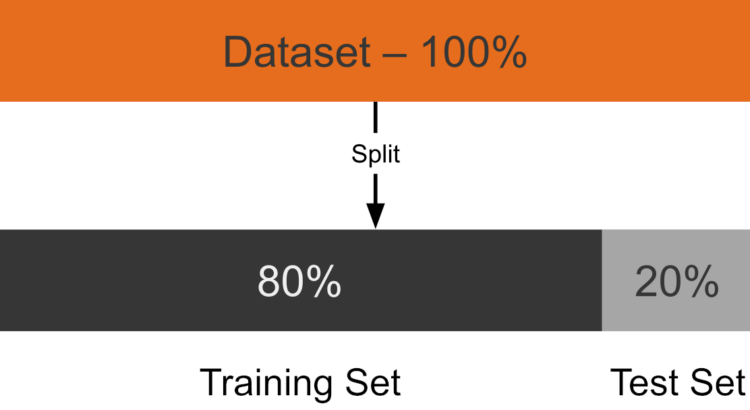
Training Models with MLJ.jl
MLJ.jl provides a uniform interface for every model family in Julia. The same machine -> fit! -> predict pattern works regardless of algorithm, making it straightforward to swap models or run comparisons.
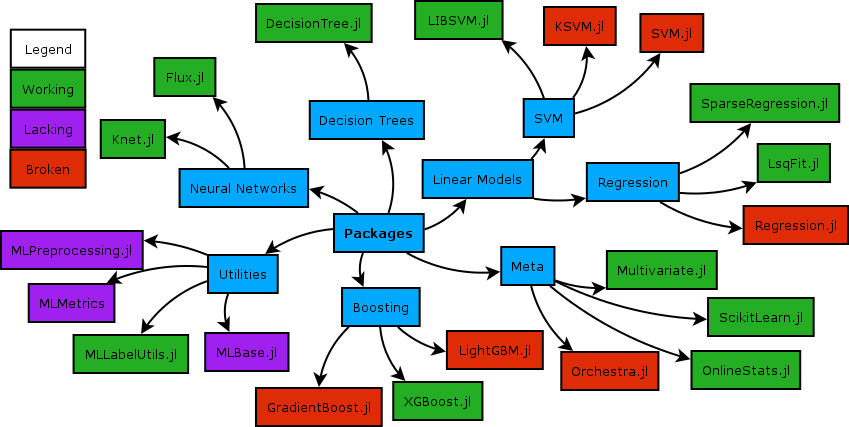
Three models are trained and compared. The evaluation metric is AUC (Area Under the ROC Curve): 1.0 is a perfect classifier, 0.5 is random guessing.
using ROCAnalysis
function evaluate_model(model, X_train, y_train, X_test, y_test)
mach = machine(model, X_train, y_train)
fit!(mach; verbosity = 0)
# pdf(..., "1") extracts the predicted probability of the positive class
probs = pdf.(predict(mach, X_test), "1")
score = auc(roc(probs, y_test .== "1"))
return score, mach
endLogistic Regression with L1 Regularization
Logistic regression is the transparent baseline. L1 regularization (Lasso) drives the coefficients of irrelevant features exactly to zero, effectively selecting the most informative predictors and keeping the model interpretable.
See: MLJLinearModels.jl documentation
using MLJLinearModels
logreg = MLJLinearModels.LogisticClassifier(penalty = :l1, lambda = 0.0428)
auc_lr, mach_lr = evaluate_model(logreg, X_train, y_train, X_test, y_test)
println("Logistic Regression AUC: $auc_lr")Random Forest
Random forests build an ensemble of decision trees on random subsets of the training data and average their predictions. They handle non-linear relationships naturally, require minimal preprocessing, and are robust to correlated and noisy features.
See: MLJDecisionTreeInterface.jl documentation
using MLJDecisionTreeInterface
rf = MLJDecisionTreeInterface.RandomForestClassifier(n_trees = 100, max_depth = 10)
auc_rf, mach_rf = evaluate_model(rf, X_train, y_train, X_test, y_test)
println("Random Forest AUC: $auc_rf")n_trees = 100 provides a stable ensemble; max_depth = 10 caps each tree's complexity to reduce overfitting.
XGBoost
XGBoost (eXtreme Gradient Boosting) trains trees sequentially, with each tree correcting the residual errors of the previous one. It consistently delivers strong performance on structured tabular data and is often the best-performing algorithm in clinical prediction benchmarks.
See: MLJXGBoostInterface.jl documentation
using MLJXGBoostInterface
xgb = MLJXGBoostInterface.XGBoostClassifier(num_round = 100, max_depth = 5, eta = 0.1)
auc_xgb, mach_xgb = evaluate_model(xgb, X_train, y_train, X_test, y_test)
println("XGBoost AUC: $auc_xgb")eta = 0.1 is the learning rate - how much each tree corrects the model. Lower values generalise better but need more boosting rounds. max_depth = 5 limits tree depth to control model complexity.
Full Pipeline Summary
The complete PLP workflow from raw database to evaluated model:
| Step | Script | Key Package |
|---|---|---|
| Study initialization | setup | HealthBase.jl |
| Cohort download | setup | OHDSIAPI.jl |
| Data exploration | src/01_data_loader.jl | DuckDB.jl + PrettyTables.jl |
| Cohort SQL translation & execution | src/02_cohort_definition.jl | OHDSICohortExpressions.jl + FunSQL.jl |
| Feature extraction | src/03_feature_extraction.jl | DuckDB.jl + DataFrames.jl |
| Distribution check | src/04_distribution_check.jl | Statistics |
| Outcome labeling | src/05_outcome_attach.jl | DataFrames.jl |
| Imputation, standardization, encoding | src/06_preprocessing.jl | CategoricalArrays.jl + Statistics |
| Train / test split | src/06_preprocessing.jl | MLJ.jl |
| Model training & evaluation | src/07_train_model.jl | MLJLinearModels · MLJDecisionTreeInterface · MLJXGBoostInterface |
| AUC scoring | src/07_train_model.jl | ROCAnalysis.jl |
The entire workflow is orchestrated by run.jl:
# run.jl
steps = [
("Defining cohorts", joinpath("src", "02_cohort_definition.jl")),
("Extracting features", joinpath("src", "03_feature_extraction.jl")),
("Checking distributions", joinpath("src", "04_distribution_check.jl")),
("Attaching outcomes", joinpath("src", "05_outcome_attach.jl")),
("Preprocessing", joinpath("src", "06_preprocessing.jl")),
("Training models", joinpath("src", "07_train_model.jl")),
]
for (label, path) in steps
println("\n── $label ────")
include(path)
endTo reproduce the full pipeline:
julia --project=. run.jlReferences
Reps, J. M., Schuemie, M. J., Suchard, M. A., Ryan, P. B., & Rijnbeek, P. R. (2018). Design and implementation of a standardized framework to generate and evaluate patient-level prediction models using observational healthcare data. Journal of the American Medical Informatics Association, 25(8), 969–975. https://doi.org/10.1093/jamia/ocy032