JuliaHealth Ecosystem Audit: Documentation, CI and Package Health
Introduction
Hi everyone! I’m Kosuri Lakshmi Indu, a final-year undergraduate student majoring in Computer Science and a Google Summer of Code 2025 contributor. Over the past year, I’ve had the wonderful opportunity to work with the JuliaHealth community, where I’ve been learning, contributing and getting involved in various projects focused on improving how we work with healthcare data and strengthen our ecosystem.
This blog post is part of my work on a NumFOCUS Small Development Grant project focused on Improving JuliaHealth Documentation Accessibility for Community Onboarding. The grant supports three main goals:
- Attracting new community members and contributors: by making documentation more accessible and centralized
- Highlighting JuliaHealth workflows: through practical guides and examples
- Strengthening community robustness: by improving documentation practices, CI pipelines and package maintenance
Why This Audit?
As the first phase of this grant work, I conducted a comprehensive audit of the entire JuliaHealth ecosystem. The goal was simple: understand the current state of our org packages so we can identify what’s working well and where we need help.
The JuliaHealth organization currently maintains 60 (48 packages + 12 non-package) repositories focused on healthcare, medical imaging, bioinformatics and health data analysis. But without a systematic way to assess all packages, it’s impossible to know which ones are well-documented, which have good testing infrastructure and which are actively maintained. So I built an automated audit system to evaluate the entire ecosystem across a set of metrics.
This audit helps us answer important questions:
- How many packages have documentation deployed for users?
- Do packages have contributing guidelines to help newcomers get started?
- Which packages track code coverage to ensure test quality?
- Are packages actively maintained or have some gone quiet?
- Who are the active maintainers across the ecosystem?
In this blog post, I’ll walk you through what I discovered about the JuliaHealth ecosystem. The work is well documented in repository: JuliaHealthAudit
General Registry Status
The General Registry is Julia’s central package repository that makes packages easily discoverable and installable via the built-in package manager. Being registered means users can simply run using PackageName or ] add PackageName without needing to know the GitHub URL. Out of 48 packages (1 forked), 33 are registered in the General Registry, while 14 remain unregistered.
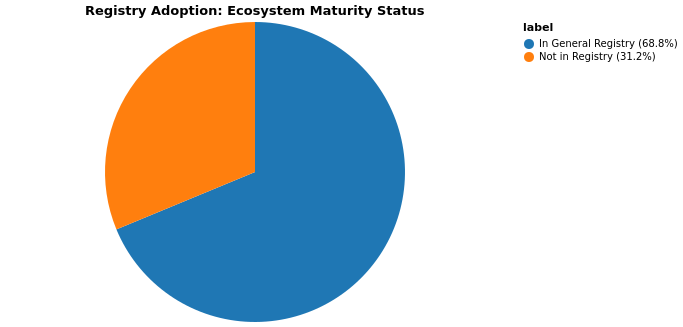
Packages NOT in General Registry:
- CTakesParser.jl
- CloToP.jl
- GitHubAnalytics.jl
- HealthDash.jl
- HealthLLM.jl
- IPUMS.jl
- ITKIOWrapp.jl
- MTIWrapper.jl
- OHDSIAPI.jl
- OMOPCDMFeasibility.jl
- OMOPCDMPathways.jl
- PubMedMiner.jl
- Thunderbolt.jl
- WrapperITKIO.jl
Packages with Repo Link Mismatches (7): What I have noticed is that, even though the repo-link stated is different, it is redirecting to the JuliaHealth page.
- BlindingIndex.jl
- DICOMTree.jl
- MedEval3D.jl
- MedPipe3D.jl
- NeuroAnalyzer.jl
- OMOPVocabMapper.jl
Archived and Forked Packages
Archived packages are read-only repositories that are no longer actively maintained. Forked packages originate from other repositories and may have development happening elsewhere.
Archived Packages (2):
- WrapperITKIO.jl
- ITKIOWrapp.jl
Forked Packages (1):
- NCEI.jl
Detailed Findings
Now let’s dive deeper into specific aspects of the JuliaHealth ecosystem. This section breaks down the audit findings by categories:
- Documentation & README
- CI/CD, Testing & Coverage
- Structure, Licensing & Maturity
- Activity, Releases & Engagement
- Issues & PR Health
- Contributors (Cross-Ecosystem)
Documentation & README
Documentation is crucial for package adoption and contribution. We evaluated whether packages have a docs/ directory, use Documenter.jl, deploy to GitHub Pages, and include contributor guidelines.
Documentation Coverage
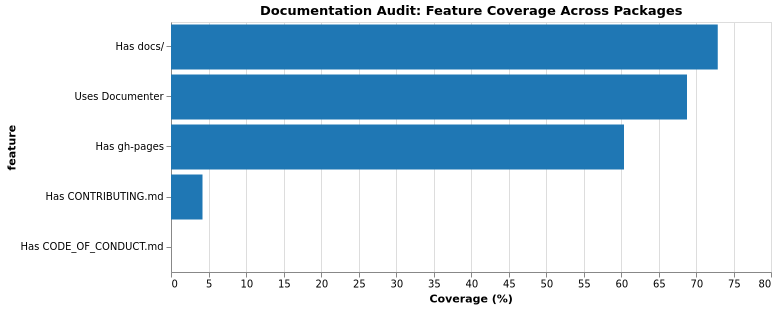
GitHub Pages Deployment
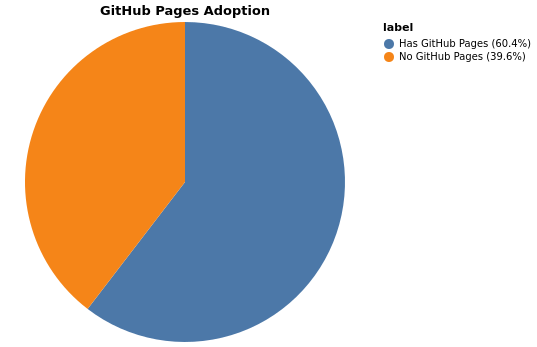
Packages without GitHub Pages:
- DICOM.jl
- NeuroAnalyzer.jl
- MedEye3d.jl
- MedPipe3D.jl
- DICOMTree.jl
- MedEval3D.jl
- HealthSampleData.jl
- ITKIOWrapper.jl
- OMOPVocabMapper.jl
- BlindingIndex.jl
- NCEI.jl
- PubMedMiner.jl
- CTakesParser.jl
- HealthDash.jl
- HealthLLM.jl
- CloToP.jl
- WrapperITKIO.jl
- ITKIOWrapp.jl
- MTIWrapper.jl
README Completeness
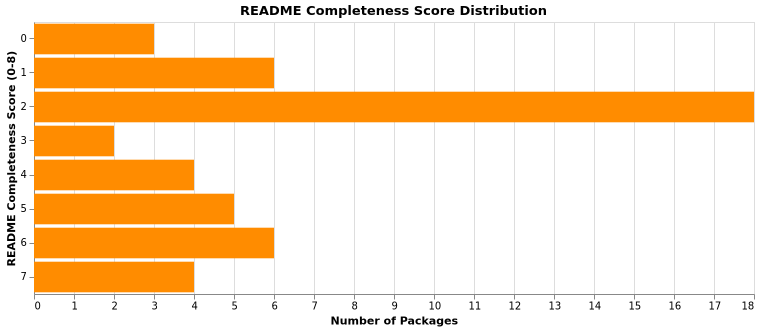
README completeness scoring (0–8), one point each:
- Install/quickstart instructions
- Usage or examples section
- Contributing guidance
- Lists that break up dense text
- Links to docs/issues/community
- Code blocks or inline examples
- Badges (build, coverage, license, etc.)
- Section headers (##) for structure
CI/CD, Testing & Coverage
Continuous Integration and testing infrastructure ensure code quality and catch bugs early. We examined CI adoption, CI status, code coverage tracking, and testing practices.
CI/CD Adoption
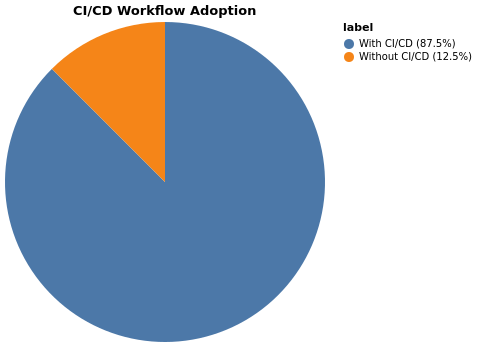
Packages missing CI workflows:
- MedPipe3D.jl
- MedEval3D.jl
- HealthDash.jl
- CloToP.jl
- WrapperITKIO.jl
- ITKIOWrapp.jl
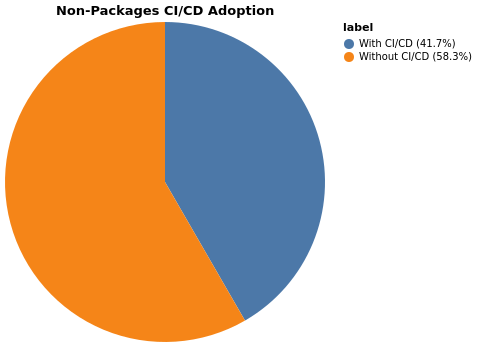
Non-package repositories missing CI workflows:
- MedPipe3DTutorial
- OMOPCDMPredictor
- ObservationalHealthSubecosystemPaper
- JuliaHealthEcosystemPaper
- JuliaHealthZoo
CI Status
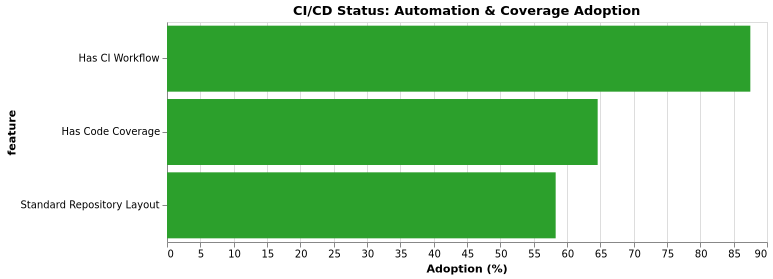
Code Coverage
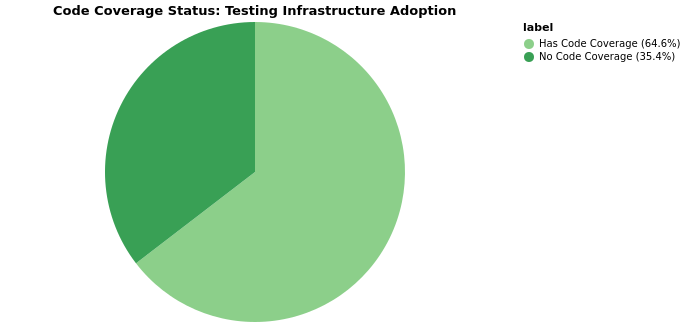
Packages without code coverage configured:
- BloodFlowTrixi.jl
- NeuroAnalyzer.jl
- BioMedQuery.jl
- MedEye3d.jl
- ARules.jl
- MedPipe3D.jl
- DICOMTree.jl
- MedEval3D.jl
- ITKIOWrapper.jl
- OMOPCDMDatabaseConnector.jl
- BlindingIndex.jl
- PubMedMiner.jl
- HealthDash.jl
- GitHubAnalytics.jl
- CloToP.jl
- WrapperITKIO.jl
- ITKIOWrapp.jl
Structure, Licensing & Maturity
Proper package structure, licensing clarity, and maturity indicators help users assess reliability. We evaluated standard layout compliance, registry status, archive status, maturity tiers, and license coverage.
For clarity, in this audit a package is considered to follow the “standard layout” when it meets the required items and all recommended checks below.
- Required items (all required):
has_src_dir: repository contains asrc/directoryhas_project_toml:Project.tomlis presenthas_license: a LICENSE file is present
- Recommended checks (all recommended):
has_test_dir:test/directory existshas_docs_dir:docs/directory existshas_ci: CI workflows exist (e.g.,.github/workflows/)uses_documenter: uses Documenter.jl for docscode_coverage_config: code coverage reporting configured (e.g., Codecov)
Packages marked as following the standard layout must satisfy every item in both lists.
Structure Compliance
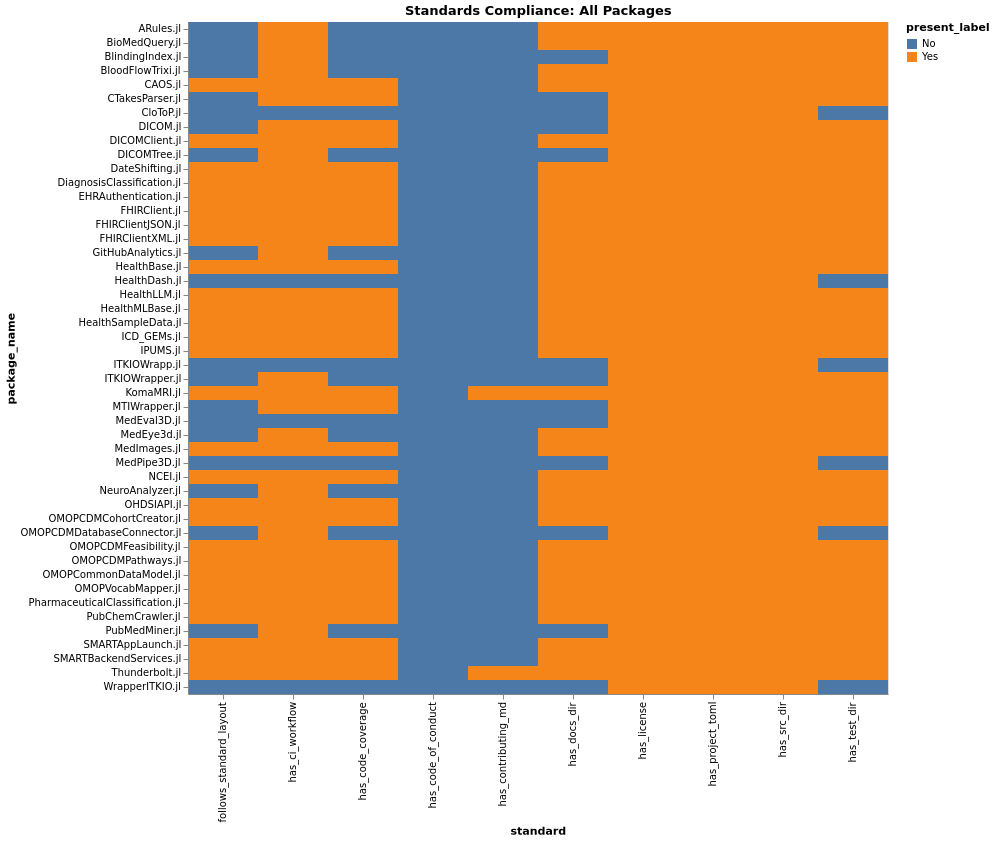
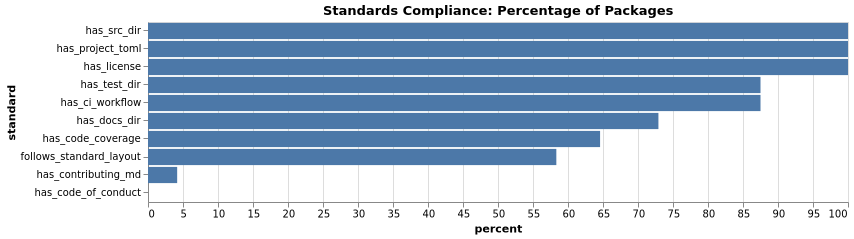
Standards Percentages
Style Guide Distribution
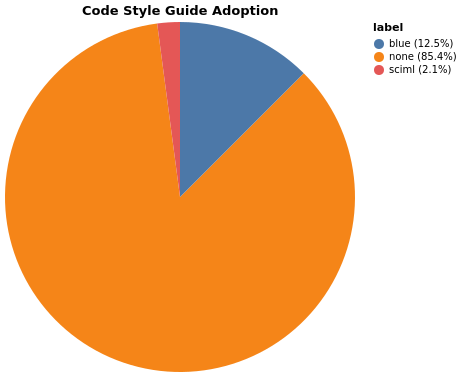
Packages Following Blue or SciML Style Guides:
| Package | Style Guide |
|---|---|
| KomaMRI.jl | Blue |
| HealthSampleData.jl | Blue |
| Thunderbolt.jl | SciML |
| OMOPCDMFeasibility.jl | Blue |
| OMOPCDMPathways.jl | Blue |
| HealthLLM.jl | Blue |
| IPUMS.jl | Blue |
Maturity Tiers
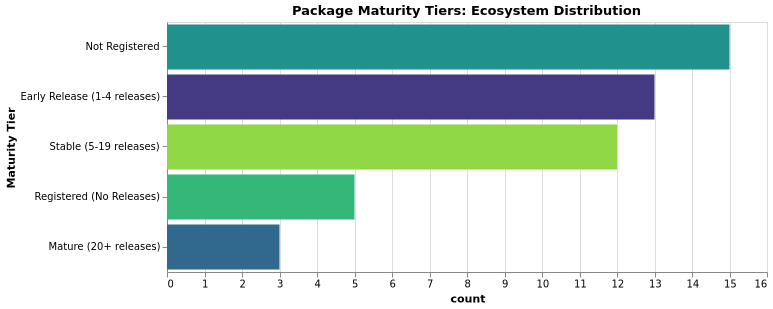
| Tier | Criteria |
|---|---|
| Mature | In General Registry; releases_count ≥ 20 |
| Stable | In General Registry; 5 ≤ releases_count ≤ 19 |
| Early Release | In General Registry; 1–4 releases |
| Registered (No Releases) | In General Registry; releases_count = 0 |
| Not Registered | Not in General Registry |
License Presence & Types
All JuliaHealth packages have licenses – 100% license compliance! This is an important ecosystem signal showing professionalism and legal clarity.
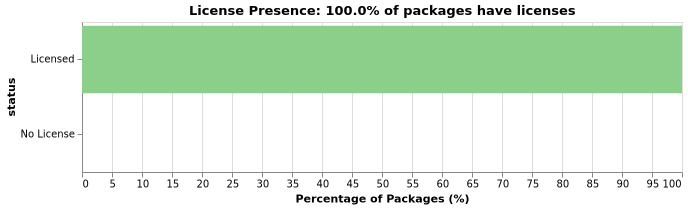
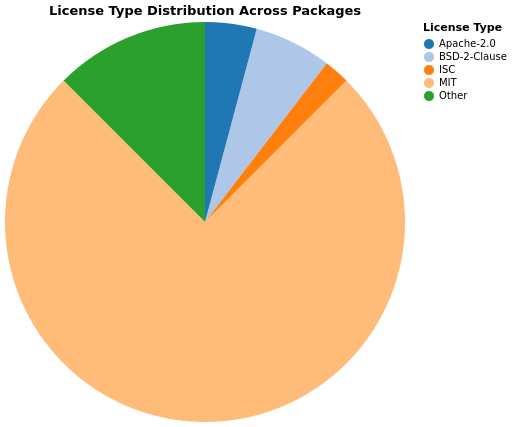
License types vary across the ecosystem. The most common licenses include MIT, Apache-2.0 and BSD variants, reflecting standard open-source practices:
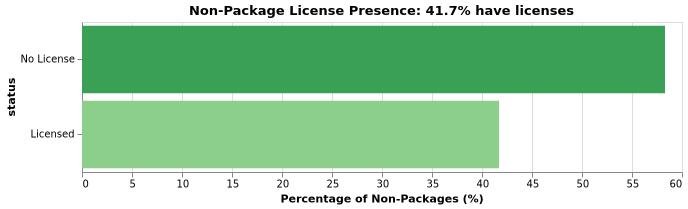
Non-Package License Coverage
Non-package repositories also show strong license compliance:
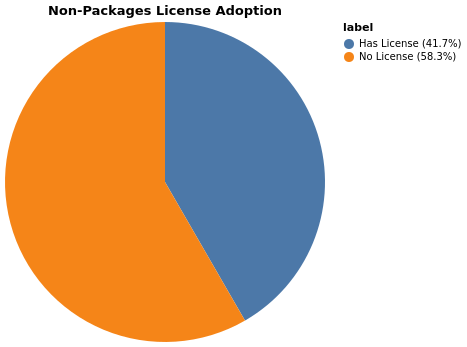
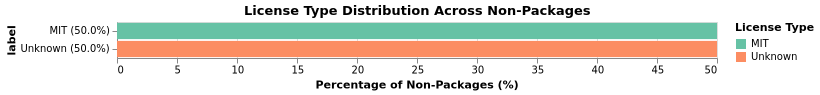
Non-package repositories missing a LICENSE file:
- JuliaHealthEcosystemPaper
- JuliaHealthWebsiteOld
- JuliaHealthZoo
- MedPipe3DTutorial
- ObservationalHealthSubecosystemPaper
Activity, Releases & Engagement
Community engagement and maintenance activity indicate package health. We analyzed activity recency, releases, stars, contributors, and popularity.
Maintainers & Maintenance Status
Understanding who maintains packages and their current activity status is critical for ecosystem health. We identify maintainers as people with push/admin/maintain access who have committed code in the last 10 years. Active maintainers are those who contributed within the last 6 months.
Maintainers Comparison (Packages):
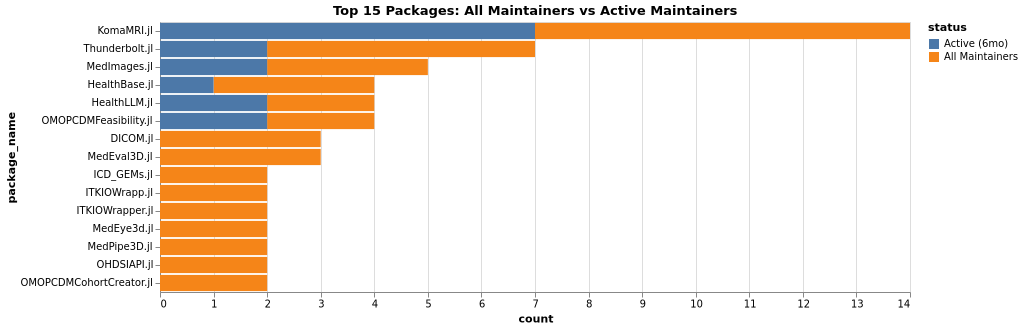
This chart shows all maintainers (people with push access who committed in the last 10 years) versus active maintainers (those who contributed in the last 6 months). The gap between the two bars indicates maintainer availability - a large gap suggests maintainers who could help but aren’t currently active.
Maintenance Status (Packages):
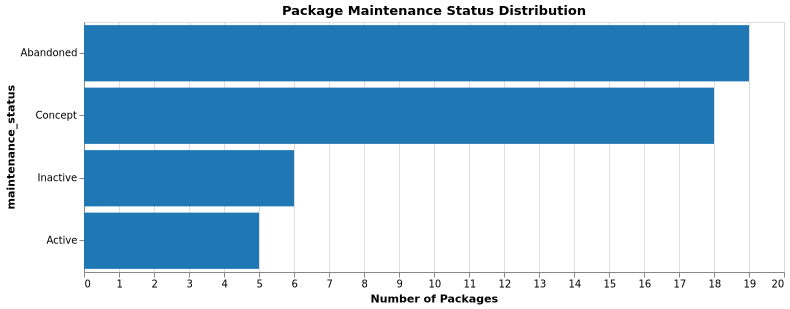
Classification criteria:
| Status | Criteria |
|---|---|
| Concept | Less than 1 release |
| Active | Has active maintainers (contributed in last 6 months) |
| Inactive | Has maintainers but no active ones, last activity < 540 days (~18 months) |
| Abandoned | No maintainers OR last activity ≥ 540 days |
Maintainers Comparison (Non-Packages):
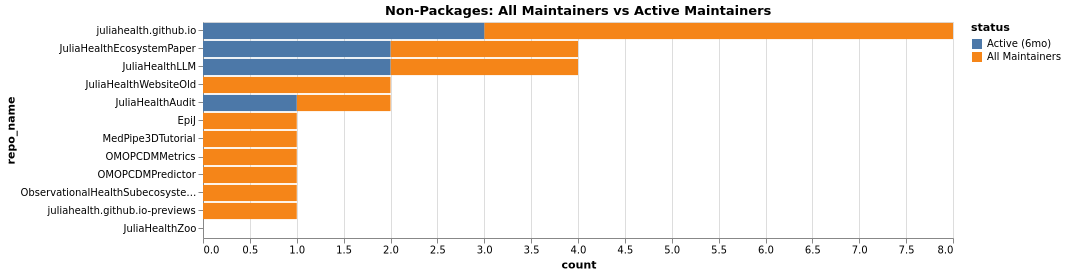
Maintenance Status (Non-Packages):
The maintenance status distribution helps identify which packages and repositories need immediate attention. Active packages have maintainers currently engaged, Inactive packages have maintainers who could be re-engaged, Abandoned packages need new maintainers or archival consideration, and Concept packages are experimental or pre-release.
Activity Recency
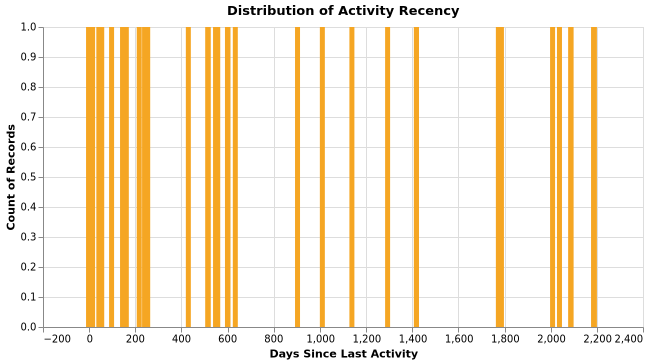
Releases Distribution
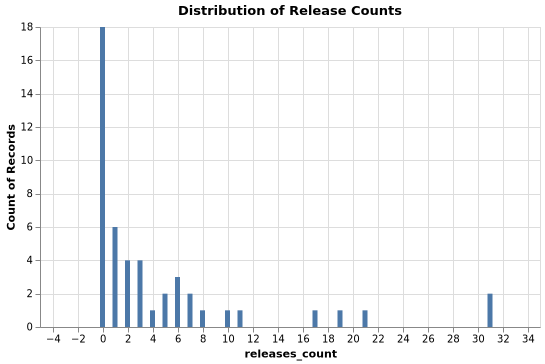
Stars Distribution
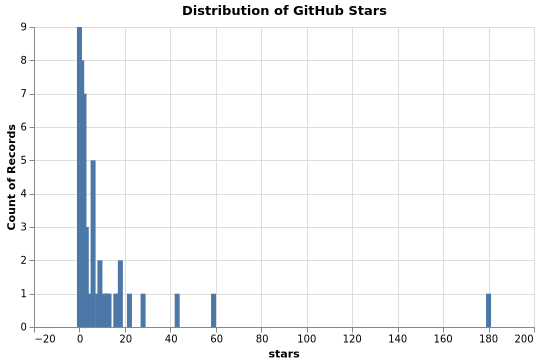
Contributors Distribution
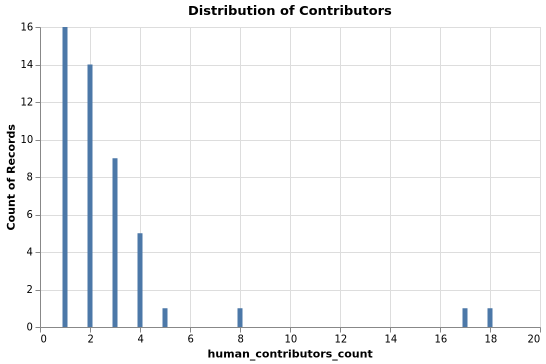
Top Packages by Stars
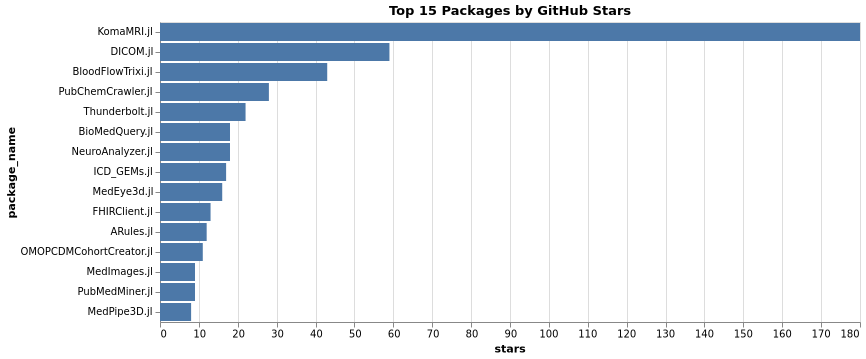
Top Packages by Monthly Downloads
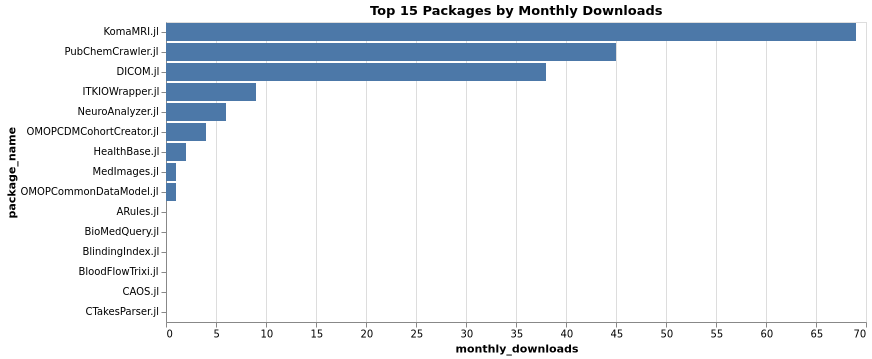
Top Packages by Total Downloads
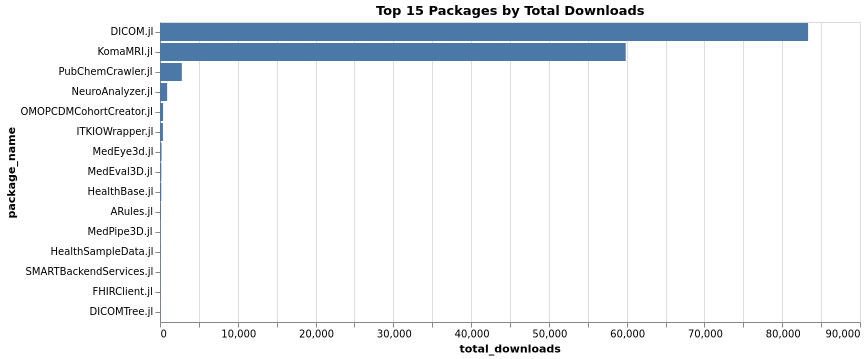
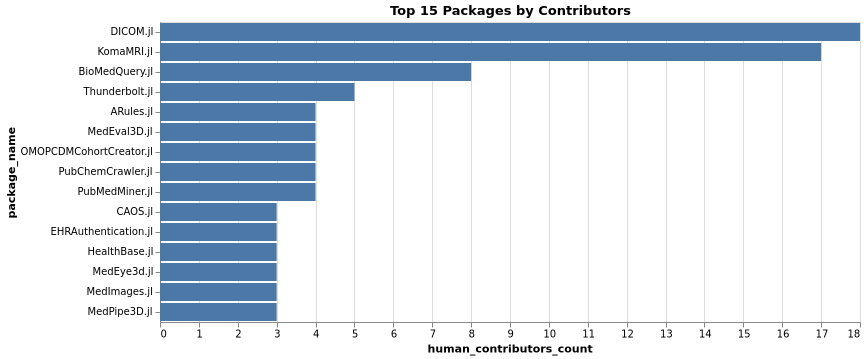
Top Packages by Contributors
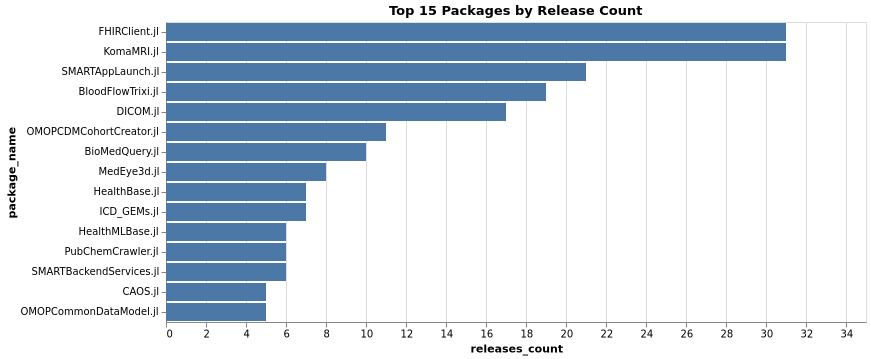
Top Packages by Releases
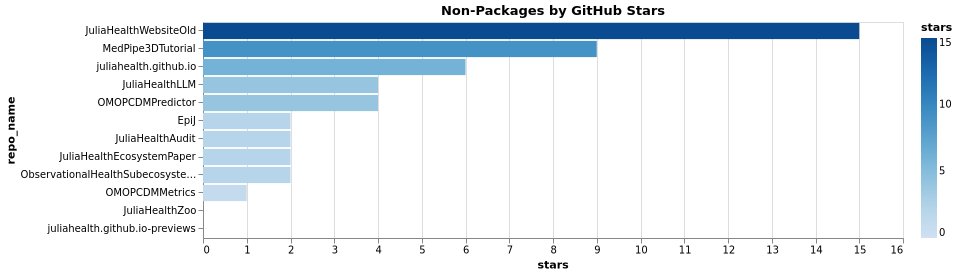
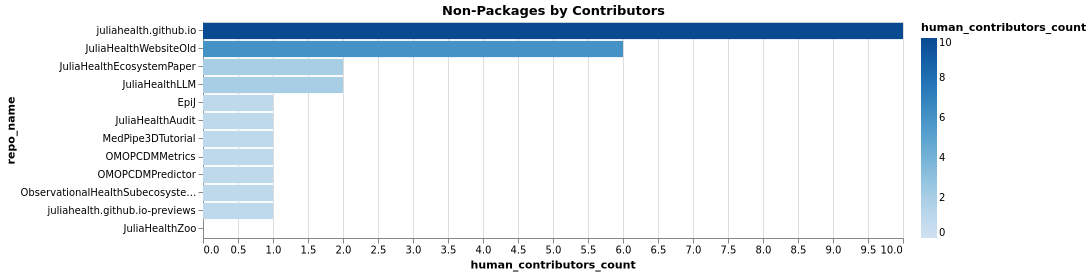
Non-Package Stars and Contributors
Issues & PR Health
Issues Comparison (Packages)
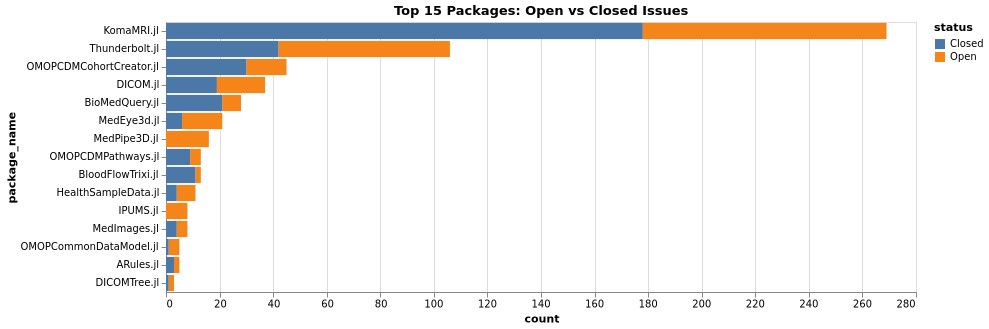
PRs Comparison (Packages)
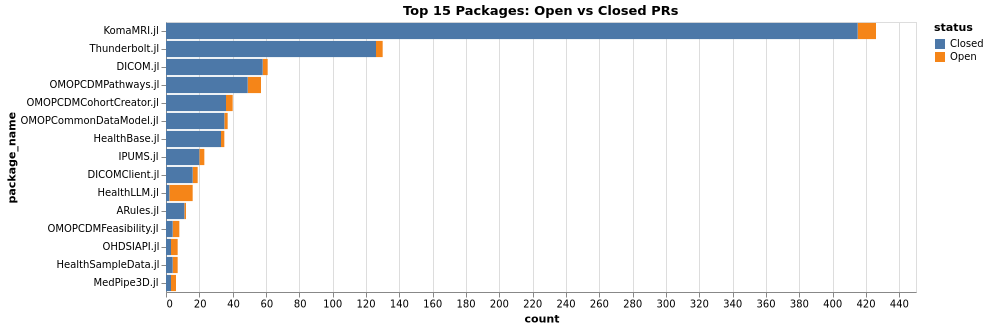
Issues Comparison (Non-Packages)
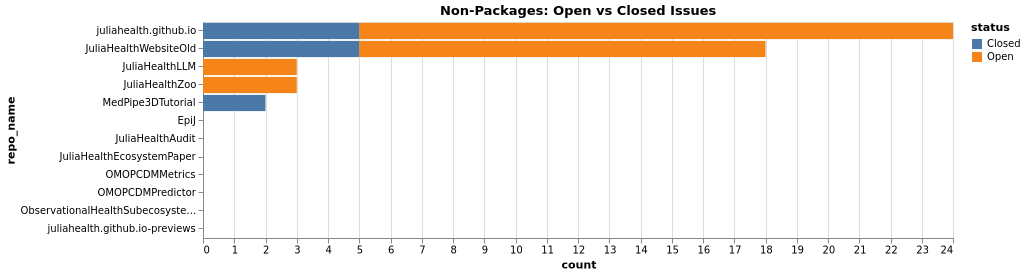
PRs Comparison (Non-Packages)
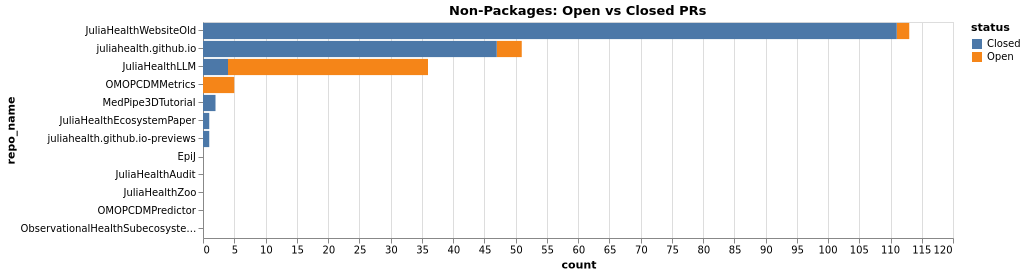
Contributors (Cross-Ecosystem)
Top Contributors by Total
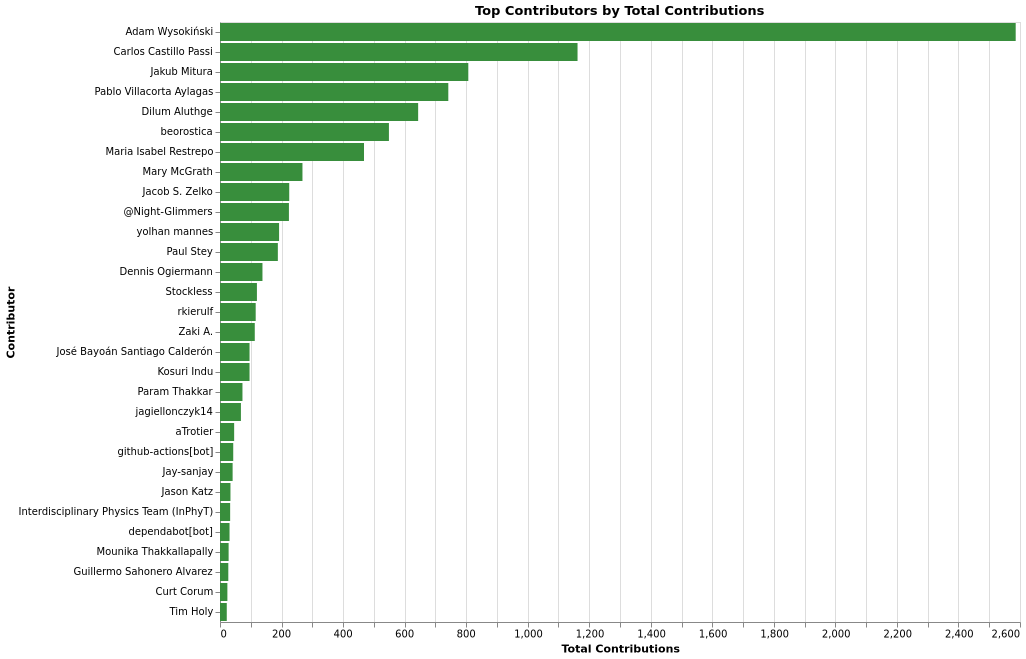
Top Contributors by Repo Count
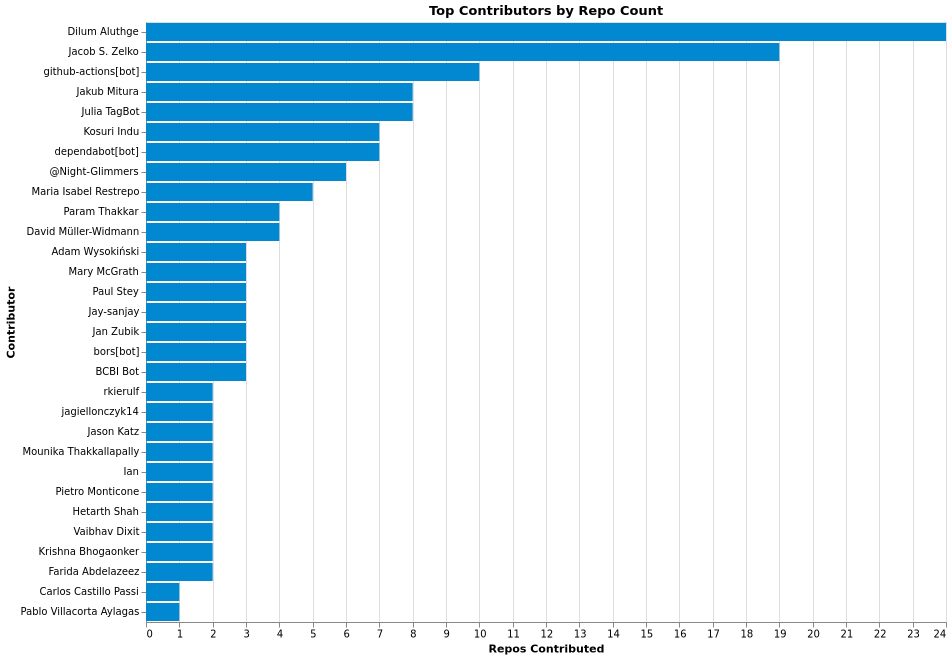
Contribution Distribution
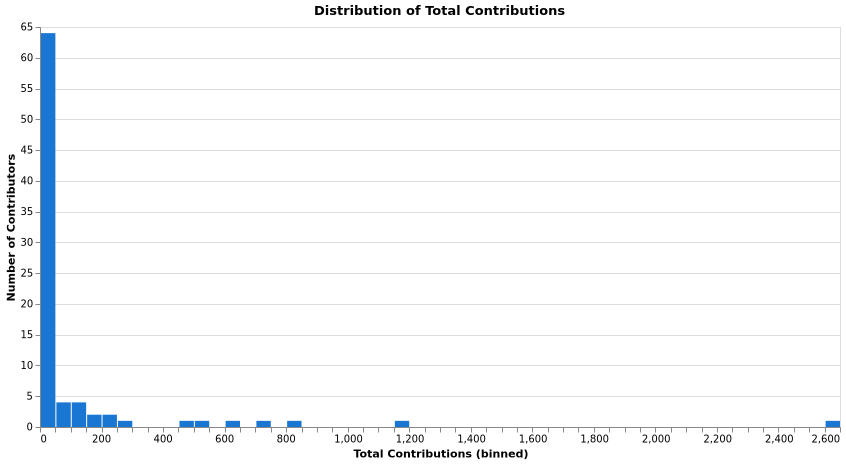
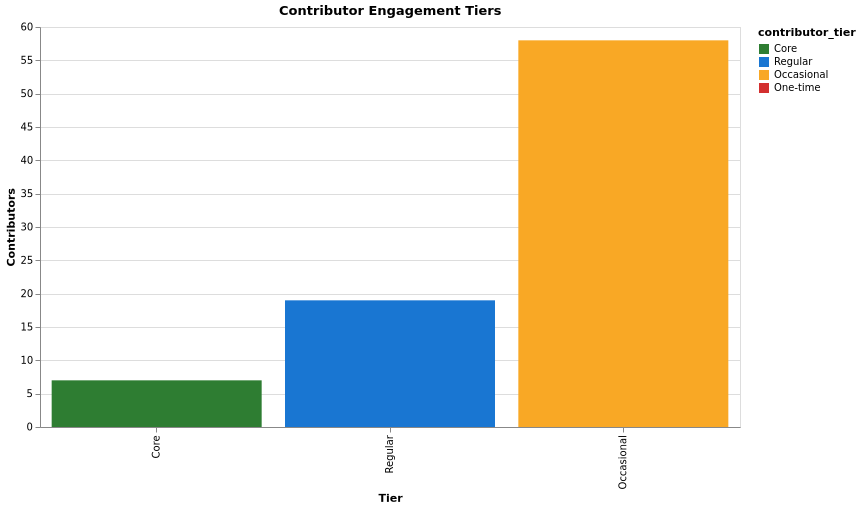
Engagement Tiers
| Tier | Rule (total_contributions, num_repos_contributed) |
|---|---|
| Core | total > 500 or repos > 10 |
| Regular | total ≥ 100 or repos ≥ 3 |
| Occasional | total ≥ 10 or repos ≥ 1 |
| One-time | Otherwise |
Actionables
Looking at our findings, here are concise, prioritized actions organized in the updated categories. Each item is phrased as a small, actionable task that can be addressed in a single PR or issue.
Documentation & README
- Rationale: Good documentation makes it easy for users to learn and use the package. Good docs also help contributors get started and contribute safely.
- Actions:
- Deploy
docs/to GitHub Pages. - Add a short
CONTRIBUTING.mdand aCODE_OF_CONDUCT.md. - Add a well-defined example and description with a code block to the
README. - Use
Documenter.jlto build docs when appropriate.
- Deploy
CI/CD, Testing & Coverage
- Rationale: CI runs tests automatically to find bugs early. Coverage helps identify parts of the code that need more tests.
- Actions:
- Add a GitHub Actions workflow that runs tests and builds docs.
- Enable coverage reporting and add a coverage badge (codecov) to the
README. - Add
test/for key functionality.
Structure, Licensing & Maturity
- Rationale: A standard layout makes packages easier to install and use. Published releases and clear licensing let others depend on stable, compliant versions. A shared code style reduces trivial review comments and confusion, and CI checks catch obvious issues before human review.
- Actions:
- Finalize a standard template for package structure.
- Publish a release and register in the General Registry if ready; fix registry metadata if needed.
- Adopt
JuliaFormatterand add formatting to CI. - Add basic lint or build checks to CI (e.g., docs build, tests pass).
Activity, Releases & Engagement
- Rationale: Clear labels and simple guides help new contributors find tasks. Public maintainer info makes it easier to coordinate work and handoffs. Active maintenance signals health and reliability to users.
- Actions:
- Add
good first issueandhelp wantedlabels and link them toCONTRIBUTING.md. - Add a
MAINTAINERS.mdor contact/triage instructions in theREADME. - For Abandoned packages: Consider archiving, transferring ownership, or recruiting new maintainers.
- For Inactive packages: Reach out to existing maintainers to gauge interest in returning or recruiting help.
- For Concept packages: Decide whether to develop toward first release or archive as experimental.
- Recognize and thank active maintainers publicly to sustain engagement.
- Add
Issues & PR Health
- Rationale: Well-triaged issues and reviewed PRs keep backlog healthy and contributors unblocked.
- Actions:
- Use the issue/PR comparisons above to prioritize triage and reviews.
- Apply
good first issue/help wantedlabels to surface approachable work.
Contributors (Cross-Ecosystem)
- Rationale: Recognizing and engaging active contributors helps sustain momentum across repositories.
- Actions:
- Use the generated lists in data/results/lists to pick concrete targets and open issues or PRs against the repositories listed.
Conclusion
The JuliaHealth ecosystem is growing and has a strong foundation. Most packages follow good practices like CI/CD automation and General Registry registration. We have excellent examples like KomaMRI.jl that show what well-maintained packages look like. But there’s room for improvement. Some packages lack documentation, contributing guidelines and code of conduct files. Some valuable packages have become inactive and need new maintainers.
The good news is that every gap we identified is an opportunity to contribute. This audit is just the beginning. The goal is to make JuliaHealth more accessible so new members can easily discover packages, understand how to use them, and contribute improvements.
If you want to get involved, check out the audit repository, browse the detailed findings and pick something you’d like to work on. The JuliaHealth community is friendly and supportive. Reach out on Julia Slack (#health-and-medicine) or Discourse if you have questions. Let’s work together to make JuliaHealth even better ❤️.
Acknowledgements
This work was made possible by the NumFOCUS Small Development Grant program. Thank you to NumFOCUS for supporting ecosystem health and community sustainability. Special thanks to my mentors: Carlos Castillo Passi and Jacob Zelko (TheCedarPrince) - for project guidance, technical mentorship and insights into the JuliaHealth ecosystem
Thank you to:
- The JuliaHealth community - Over 300 members across Slack, Zulip, and Discourse who make this ecosystem welcoming and collaborative
- All package maintainers and developers - For building and maintaining the tools that make this audit possible
- The Julia community - For creating an amazing language and ecosystem
Note: This blog post was drafted with the assistance of LLM technologies to support grammar, clarity and structure.
Citation
@online{indu2026,
author = {Indu, Kosuri},
title = {JuliaHealth {Ecosystem} {Audit:} {Documentation,} {CI} and
{Package} {Health}},
date = {2026-01-23},
langid = {en}
}