PubMed
Utility functions to parse and store PubMed searches via BioServices.EUtils
Import Module
using BioMedQuery.PubMed
This module provides utility functions to parse, store and export queries to PubMed via the NCBI EUtils and its julia interface BioServices.EUtils. For many purposes you may interact with the higher level pipelines in [BioMedQuery.Processes]. Here, some of the lower level functions are discussed in case you need to assemble different pipelines.
Basics of searching PubMed
We are often interseted in searching PubMed for all articles related to a search term, and possibly restricted by other search criteria. To do so we use BioServices.EUtils. A basic example of how we may use the functions esearch and efetch to accomplish such task is illustrated below.
using BioServices.EUtils using XMLDict search_term = "obstructive sleep apnea[MeSH Major Topic]" #esearch esearch_response = esearch(db="pubmed", term = search_term, retstart = 0, retmax = 20, tool ="BioJulia") #convert xml to dictionary esearch_dict = parse_xml(String(esearch_response.data)) #convert id's to a array of numbers ids = [parse(Int64, id_node) for id_node in esearch_dict["IdList"]["Id"]] #efetch efetch_response = efetch(db = "pubmed", tool = "BioJulia", retmode = "xml", rettype = "null", id = ids) #convert xml to dictionary efetch_dict = parse_xml(String(efetch_response.data))
Handling XML responses
Many APIs return responses in XML form.
To parse an XML to a Julia dictionary we can use the XMLDict package
using XMLDict dict = parse_xml(String(response.data))
You can save directly the XML String to file using LightXML
xdoc = parse_string(esearch) save_file(xdoc, "./file.xml")
Save eseach/efetch responses
Save PMIDs to MySQL
If we are only interseted in saving a list of PMIDs associated with a query, we can do so as follows
dbname = "entrez_test" host = "127.0.0.1"; user = "root" pwd = "" #Collect PMIDs from esearch result ids = Array{Int64,1}() for id_node in esearch_dict["IdList"]["Id"] push!(ids, parse(Int64, id_node)) end # Initialize or connect to database const conn = DBUtils.init_mysql_database(host, user, pwd, dbname) # Create `article` table to store pmids PubMed.create_pmid_table!(conn) #Save pmids PubMed.save_pmids!(conn, ids) #query the article table to explore list of pmids all_pmids = BioMedQuery.PubMed.all_pmids(conn)
Export efetch response as EndNote citation file
We can export the information returned by efetch as and EndNote/BibTex library file
citation = PubMed.CitationOutput("endnote", "./citations_temp.endnote", true) nsucceses = PubMed.save_efetch!(citation, efetch_dict, verbose)
Save efetch response to MySQL database
Save the information returned by efetch to a MySQL database
dbname = "efetch_test" host = "127.0.0.1"; user = "root" pwd = "" # Save results of efetch to database const conn = DBUtils.init_mysql_database(host, user, pwd, dbname) PubMed.create_tables!(conn) PubMed.save_efetch!(conn, efetch_dict)
Save efetch response to SQLite database
Save the information returned by efetch to a MySQL database
db_path = "./test_db.db" const conn = SQLite.DB(db_path) PubMed.create_tables!(conn) PubMed.save_efetch!(conn, efetch_dict)
Exploring output databases
The following schema has been used to store the results. If you are interested in having this module store additional fields, feel free to open an issue
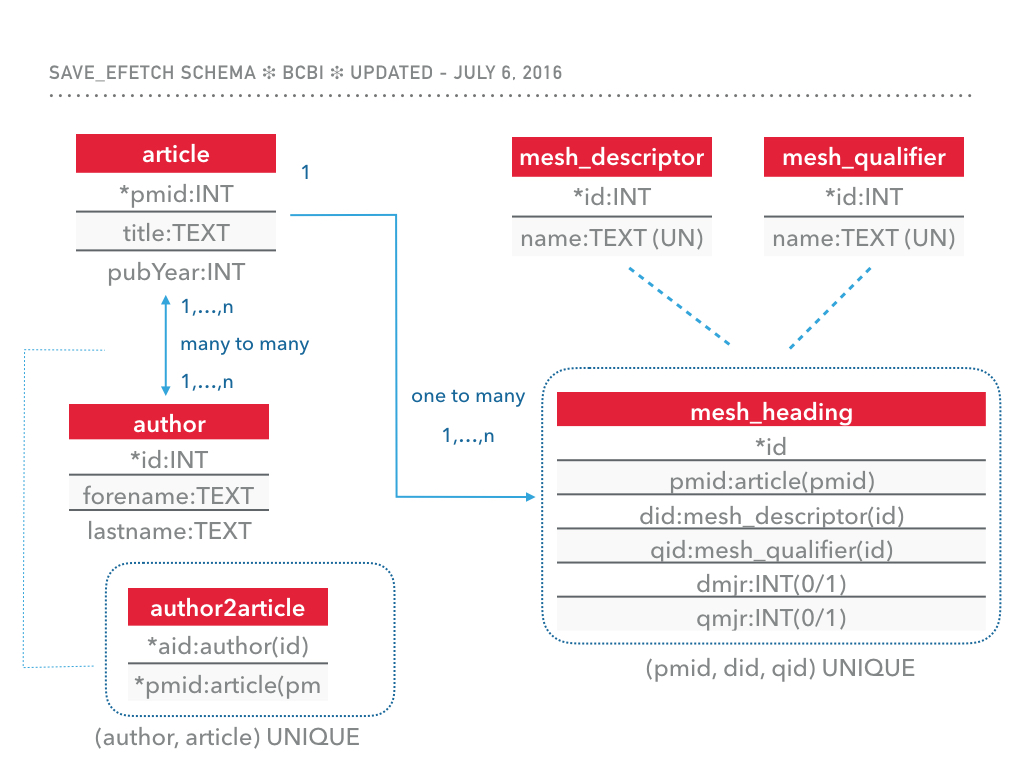
We can als eexplore the tables using BioMedQuery.DBUtils, e,g
tables = ["author", "author2article", "mesh_descriptor", "mesh_qualifier", "mesh_heading"] for t in tables query_str = "SELECT * FROM "*t*" LIMIT 10;" q = DBUtils.db_query(db, query_str) println(q) end
Index
BioMedQuery.PubMed.abstractsBioMedQuery.PubMed.abstracts_by_yearBioMedQuery.PubMed.all_meshBioMedQuery.PubMed.all_pmidsBioMedQuery.PubMed.citations_bibtexBioMedQuery.PubMed.citations_endnoteBioMedQuery.PubMed.create_pmid_table!BioMedQuery.PubMed.create_tables!BioMedQuery.PubMed.get_article_meshBioMedQuery.PubMed.get_article_mesh_by_conceptBioMedQuery.PubMed.save_efetch!BioMedQuery.PubMed.save_efetch!BioMedQuery.PubMed.save_pmids!
Structs and Functions
#
BioMedQuery.PubMed.abstracts — Method.
abstracts(db; local_medline=false)
Return all abstracts related to PMIDs in the database. If local_medline flag is set to true, it is assumed that db contains article table with only PMIDs and all other info is available in a (same host) medline database
#
BioMedQuery.PubMed.abstracts_by_year — Method.
abstracts_by_year(db, pub_year; local_medline=false)
Return all abstracts of article published in the given year. If local_medline flag is set to true, it is assumed that db contains article table with only PMIDs and all other info is available in a (same host) medline database
#
BioMedQuery.PubMed.all_pmids — Method.
all_pmids(db)
Return all PMIDs stored in the article table of the input database
#
BioMedQuery.PubMed.citations_bibtex — Function.
citations_bibtex(article::PubMedArticle, verbose=false)
Transforms a PubMedArticle into text corresponding to its bibtex citation
#
BioMedQuery.PubMed.citations_endnote — Function.
citations_endnote(article::PubMedArticle, verbose=false)
Transforms a PubMedArticle into text corresponding to its endnote citation
#
BioMedQuery.PubMed.create_pmid_table! — Method.
init_pmid_db!(conn; tablename="article")
Creates a database, using either MySQL of SQLite, with all necessary tables to store Entrez related searches. All tables are empty at this point
#
BioMedQuery.PubMed.create_tables! — Method.
create_tables!(conn)
Create and initialize tables to save results from an Entrez/PubMed search. Caution, all related tables are dropped if they exist
#
BioMedQuery.PubMed.get_article_mesh — Method.
get_article_mesh(db, pmid)
Get the all mesh-descriptors associated with a give article
#
BioMedQuery.PubMed.get_article_mesh_by_concept — Method.
get_article_mesh_by_concept(db, pmid, umls_concepts...; local_medline)
Get the all mesh-descriptors associated with a give article
Argumets:
- query_string: "" - assumes full set of results were saved by BioMedQuery directly from XML
#
BioMedQuery.PubMed.save_efetch! — Function.
pubmed_save_efetch(efetch_dict, conn)
Save the results (dictionary) of an entrez-pubmed fetch to the input database.
#
BioMedQuery.PubMed.save_efetch! — Function.
save_efetch!(output::CitationOutput, efetch_dict, verbose=false)
Save the results of a Entrez efetch to a bibliography file, with format and file path given by output::CitationOutput
#
BioMedQuery.PubMed.save_pmids! — Function.
save_pmids!(conn, pmids::Vector{Int64}, verbose::Bool=false)
Save a list of PMIDS into input database. ###Arguments:
conn: Database connection (MySQL or SQLite)pmids: Array of PMIDsverbose: Boolean to turn on extra print statements
#
BioMedQuery.PubMed.all_mesh — Method.
all_mesh(db)
Return all PMIDs stored in the article table of the input database